TECHNOLOGY
INDUCE-seq® Platform
Why measuring breaks changes everything
Genome editing holds enormous therapeutic promise, but unintended DNA damage remains a critical barrier to safe development. Off-target effects are difficult to predict and often challenging to detect using traditional approaches.
As regulatory expectations for genome-wide assessment continue to evolve, empirical measurement of off-target activity is becoming essential.
Existing methods measure the final genetic outcome after DNA repair, limiting the ability to understand how and why off-target events occur.
±ő±·¶Ů±«°ä·ˇ-˛ő±đ±ç® directly labels and captures DNA breaks in cells, enabling genome-wide detection of editing events as they happen.
By measuring break formation rather than downstream repair outcomes, the platform provides deeper insight into: nuclease behaviour, guide RNA performance, editing kinetics, cell-specific DNA repair pathways.
This enables earlier, more informed decisions and accelerates the development of safer gene-editing therapies.
Current industry challenges with off-target analysis
Technical
Indirect measurement of editing outcomes, not break events
Noisy and inconsistent signals
PCR amplification introduces bias
Operational & scalability
Growing demand for empirical, genome-wide data
Late-stage surprises increase program risk
Low confidence drives repeated testing
Regulatory & decision risk
Fragmented workflows across multiple assays
Limited scalability in discovery
Expensive, inflexible outsourcing
A single workflow combines wet lab, sequencing and integrated bioinformatics
Cell editing &
immobilization
DNA break labeling
& library prep
Sequencing
Data analysis
Reporting

±ő±·¶Ů±«°ä·ˇ-˛ő±đ±ç® is a scalable platform technology for mapping and characterizing DNA breaks, leveraging a novel PCR-free in situ break labelling approach coupled with next-generation sequencing. It enables unbiased, genome-wide detection of DNA damage induced by any nuclease-based genome editing system.
What makes this technology different:
• PCR-free workflow eliminating amplification bias
• Cell-based, in situ break capture
• Compatible with any cell type
• Applicable to any nuclease-based editor
What this enables
Confident actionable data
Scalable across the drug discovery pipeline
In house control over critical safety data
Simultaneous on- and off-target detection in one simple workflow
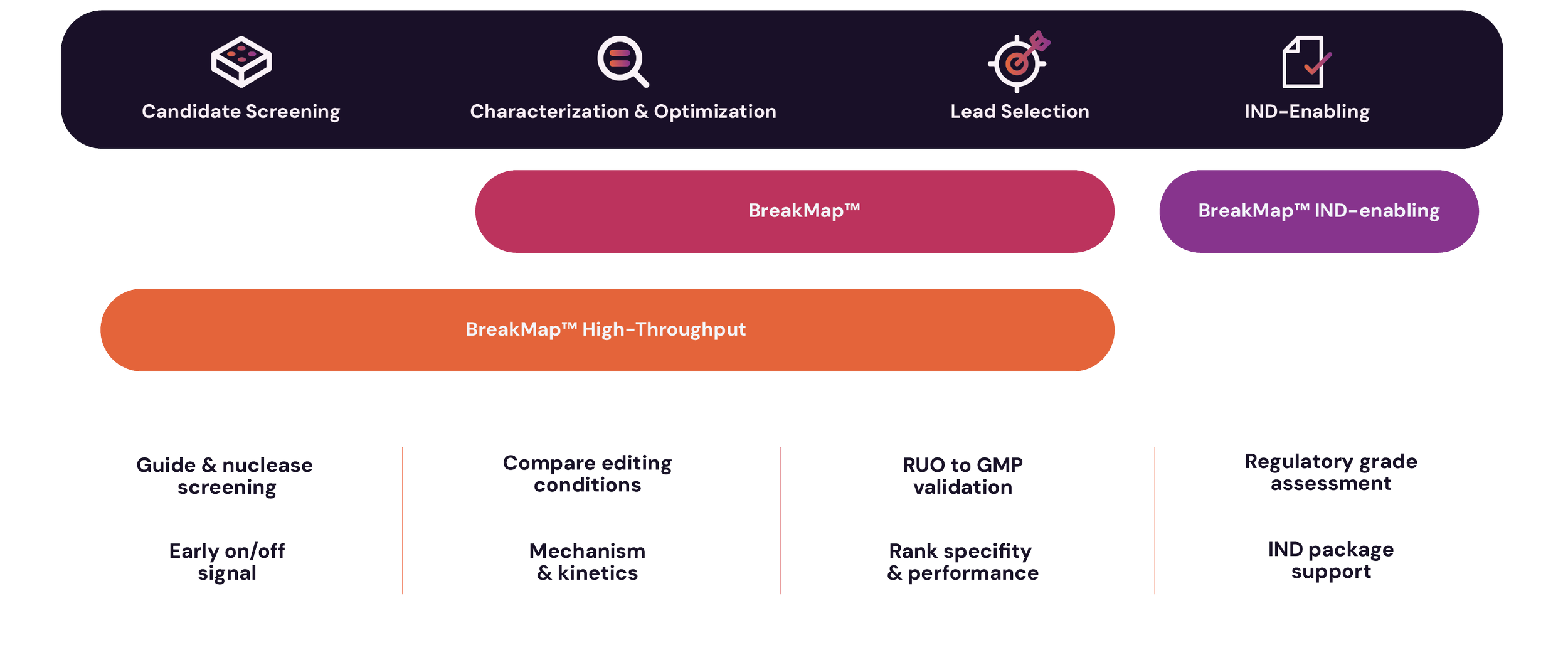
±ő±·¶Ů±«°ä·ˇ-˛ő±đ±ç® supports confident decision making from discovery to IND-enabling
Gene editing applications
INDUCE-seq® pairs its unbiased wet lab assay with an integrated bioinformatics platform designed to convert complex sequencing outputs into clear, ranked, and actionable insights facilitating:
• Off-target assessment
• Guide selection and ranking
• Editing optimization and strategy design
• On-target editing mechanism and kinetics
• Nuclease and editor characterization
All cell types: Tested examples
-
• Bone Marrow​
• Glioblastoma​
• Neurons• Motor Neurons​
​
• Lung Carcinoma​
• T Cells&˛Ô˛ú˛ő±č;​
• Dermal Fibroblasts​• Human Hepatocytes
-
• Embryonic stem cells
&˛Ô˛ú˛ő±č;​
• Hematopoietic stem and progenitor cells&˛Ô˛ú˛ő±č;​
• Induced pluripotent stem cells
• Neural progenitors -
• T309 (Brain)
• T393 (Brain)
• TK6 (Blood, lymphoblast)
• CH12F3 (Blood)
• HeLa (Uterus, epithelial)
• Jurkat (T lymphocyte)
• HEK293 (Embryonic Kidney)
• KBM7 (Bone marrow)
• MCF10A (Breast)
• MCF7 (Breast)
• RPE1 (Retina)
• SH-SY5Y (Neuroblastoma)
• RPE1 (Retina)
• A549 (Lung)
• U2OS (Bone)
• HepG2 (Liver)
• CX18 (Neuronal, Brain)
• GM24385 (B-Lymphocyte)